library(dplyr)
library(ggplot2)
library(tibble)
library(tidyr)
set.seed(123)Introduction
Pre-post analysis is a common statistical approach used to evaluate the effectiveness of an intervention or treatment by comparing measurements taken before and after the intervention. This type of analysis is particularly useful in fields such as medicine, psychology, education, and social sciences, where researchers aim to assess changes in outcomes resulting from specific interventions.
There are two common statistical approaches used for pre-post analysis:
- Repeated Measures Analysis of Variance (RM-ANOVA)
- Analysis of Covariance (ANCOVA)
RM-ANOVA is used when the same participants are measured multiple times, allowing researchers to account for within-subject variability. ANCOVA, on the other hand, is used to control for potential confounding variables by including them as covariates in the analysis. In this context, the ANCOVA model includes the pre-intervention scores as a covariate. Both models are based on linear regression under the hood, but they differ in how they handle the data structure and the assumptions they make.
Data
We will simulate data for a pre-post study with three groups (for example, two treatment groups and a control group). Each group will have 100 participants, and we will generate pre- and post-intervention scores by drawing from a bivariate normal distribution with specified means, standard deviations, and correlations.
First, we import packages and initialize the random number generator for reproducibility:
Next, we define the group size:
n = 100Now, we define the group-specific means for pre- and post-intervention scores:
(means = matrix(
c(51, 49, 50, 55.5, 53, 52),
nrow=3,
dimnames=list(NULL, c("pre", "post"))
)) pre post
[1,] 51 55.5
[2,] 49 53.0
[3,] 50 52.0We also define a standard deviation, which we assume to be identical for pre- and post-scores:
s = 10Finally, we define a correlation between pre- and post-scores:
rho = 0.7Using these parameters, we create a covariance matrix for the simulated data:
(sigma = matrix(c(s^2, rho * s^2, rho * s^2, s^2), nrow=2)) [,1] [,2]
[1,] 100 70
[2,] 70 100We then create a helper function to generate multivariate normal data:
rmvnorm = function(n, mu, sigma) {
z = matrix(rnorm(n * length(mu)), nrow=n)
z %*% chol(sigma) + matrix(mu, nrow=n, ncol=length(mu), byrow=TRUE)
}Using this function, we generate the simulated data for all three groups:
m = do.call(rbind, lapply(1:nrow(means), function(i) rmvnorm(n, means[i, ], sigma)))
df = tibble(
id=as.character(1:(3*n)),
group=factor(rep(c("A", "B", "C"), each=n)),
pre=m[, 1],
post=m[, 2]
) |>
pivot_longer(cols=c("pre", "post"), names_to="time", values_to="score")
df[["time"]] = factor(df[["time"]], levels=c("pre", "post"))Here’s a scatter plot showing the three groups pre- and post-intervention:
means_data = df |>
group_by(group, time) |>
summarize(score=mean(score), .groups="drop")
ggplot(data=df, mapping=aes(x=group, y=score, color=time, fill=time)) +
geom_point(
position=position_dodge(width=0.5),
shape=19,
size=2,
alpha=0.5
) +
geom_point(
data=means_data,
position=position_dodge(width=0.5),
shape=23,
size=3,
color="black"
) +
geom_text(
data=means_data,
mapping=aes(label=round(score, 1)),
position=position_dodge(width=0.5),
hjust=-0.4,
size=3,
color="black"
) +
labs(x="Group", y="Score", color="Time", fill="Time") +
theme_minimal()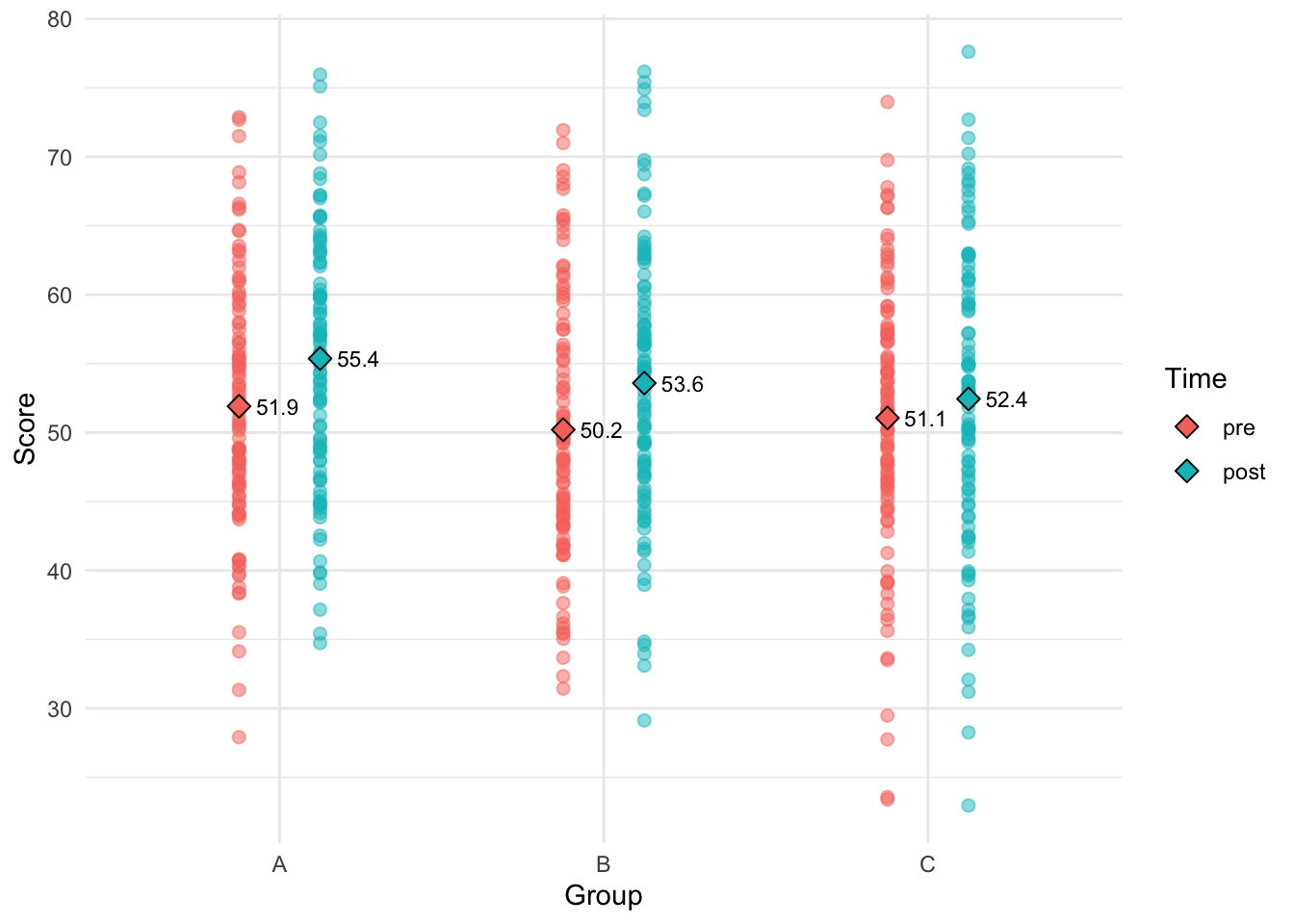
RM-ANOVA
We now analyze the data using an RM-ANOVA model with a within-subject factor time (pre vs. post) and a between-subject factor group (A, B, C):
model1 = aov(score ~ group * time + Error(id/time), data=df)
summary(model1)
Error: id
Df Sum Sq Mean Sq F value Pr(>F)
group 2 440 220.1 1.38 0.253
Residuals 297 47388 159.6
Error: id:time
Df Sum Sq Mean Sq F value Pr(>F)
time 1 1126 1126.4 40.316 8.08e-10 ***
group:time 2 139 69.4 2.485 0.085 .
Residuals 297 8298 27.9
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1We are particularly interested in the group × time interaction, which indicates whether the change from pre to post differs across groups.
ANCOVA
Another common approach for analyzing pre-post data is ANCOVA, where the post-intervention score is modeled as a function of the group and the pre-intervention score as a covariate. This approach adjusts for any baseline differences in pre-scores across groups. Note that we need to expand the time column into separate pre and post columns for this analysis:
df = pivot_wider(df, names_from=time, values_from=score)
model2 = lm(post ~ group + pre, data=df)
anova(model2)Analysis of Variance Table
Response: post
Df Sum Sq Mean Sq F value Pr(>F)
group 2 434.8 217.4 4.4076 0.01299 *
pre 1 14215.6 14215.6 288.2365 < 2e-16 ***
Residuals 296 14598.5 49.3
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1In this model, we are interested in the effect of group on the post-intervention score.
Furthermore, the additive model assumes equal regression slopes (no group × pre interaction), so we need to check this assumption:
anova(lm(post ~ group * pre, data=df))Analysis of Variance Table
Response: post
Df Sum Sq Mean Sq F value Pr(>F)
group 2 434.8 217.4 4.4363 0.01264 *
pre 1 14215.6 14215.6 290.1126 < 2e-16 ***
group:pre 2 192.4 96.2 1.9634 0.14222
Residuals 294 14406.1 49.0
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1